Stanford COVID Dashboards
Wastewater Analysis
This page provides regular updates on the analysis of wastewater collections from Palo Alto in Santa Clara County, CA.
When reviewing the graphs below, it is important to look at the overall, sustained trends, instead of measurements on any given day. Samples tend to have some variability simply due to the nature of environmental samples, but the general trends over time have proven accurate and consistent with trends in clinical cases.
As of June 1, 2024, this site displays wastewater data from Palo Alto, CA. Prior to that, the display showed analysis from campus wastewater collected from more than 150 buildings on campus by Stanford’s Department of Civil & Environmental Engineering from July 2021 through May 2024. Click here to view historical campus wastewater data.
Click here to view historical surveillance testing dashboards
The ten charts above represent wastewater concentrations for SARS-CoV-2, Influenza A, Influenza B, Respiratory Syncytial Virus (RSV), Human Metapneumovirus (HMPV), Norovirus GII, Rotavirus, Mpox, Hepatitis A, and Candida auris from daily wastewater samples collected from Palo Alto, CA. The red lines show consensus smoothing (5-sample trimmed average) of the selected pathogen concentrations normalized by PMMoV concentrations in the solids. These results are shown on a linear scale and although the target/PMMoV ratio is unitless, it is multiplied by 1 million on the y axis of these plots to make the numbers easier to interpret.
Wastewater analysis is an increasingly important part of the university’s ongoing strategy to monitor the level of community transmission. SARS-CoV-2 (the virus that causes COVID-19) is shed in feces by infected individuals and can be measured in wastewater.
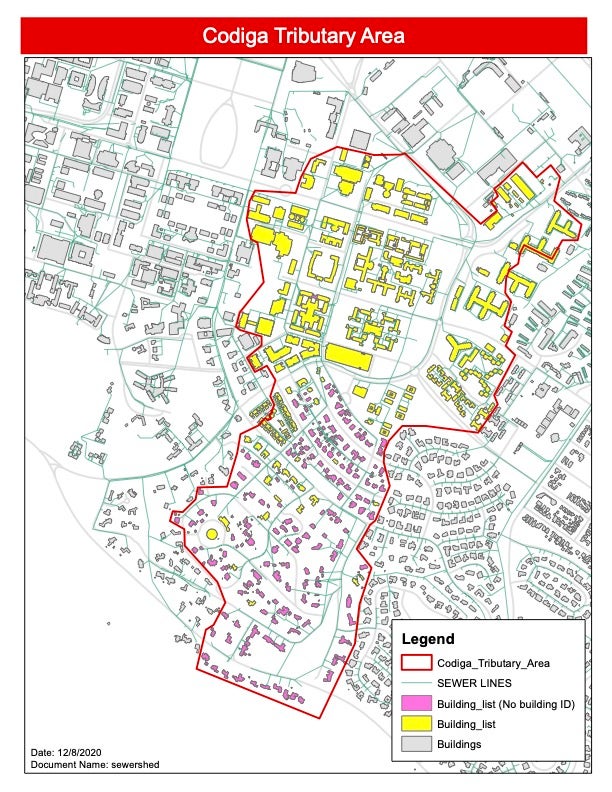
The university collects and analyzes samples from a large sewer main that drains wastewater from about 10,000 people studying, working and residing in the sewershed of Stanford’s Codiga Resource Recovery Center. This sewershed includes more than 150 buildings on campus (see map below), including faculty, staff and student housing, and classroom, research and office buildings.
Samples were collected six days a week prior to November 2022. Starting November 1, 2022, samples are collected twice a week (on Mondays and Wednesdays). Samples are analyzed within 24 hours of collection. Results are then uploaded to this dashboard.
Collection and analysis at the Stanford site began in July 2021. This effort is part of SCAN (sewer coronavirus alert network), a regional epidemiology project based at Stanford and developed in collaboration with local public health officers and 11 wastewater treatment plants in the Greater Bay Area and Central California, and the California Department of Public Health. Maps and wastewater measurements in those locations are available from SCAN.
SARS-CoV-2 Sewage Monitoring Data
Some information below is sourced from Santa Clara County Public Health.
More cases of COVID-19 in the community are associated with increased levels of SARS-CoV-2 genomes in wastewater, meaning that data from wastewater analysis can be used as an indicator of the level of transmission of COVID-19 in the community.
Wastewater analysis measures the levels of non-infectious RNA (Ribonucleic Acid) in wastewater, not the viable virus. There are no known cases of transmission resulting from exposure to wastewater.
Advantages of using wastewater to monitor COVID-19 infections
The use of wastewater to infer information about COVID-19 transmission has several potential advantages:
- Wastewater includes contributions from asymptomatic individuals and people who are unable or unwilling to obtain clinical tests, for a variety of reasons.
- It provides a mechanism to monitor the level of community transmission as clinical testing declines, and other, more convenient testing takes place (such as home-based rapid antigen tests) that are not reported to the public health departments. Wastewater information is available sooner than information from clinical testing, which means that monitoring SARS-CoV-2 in wastewater can serve as an early indicator of increasing or decreasing COVID-19 infections in the community.
- During an increase, this early information could be used to enhance public health messaging in our community to reinforce safe practices, promote more clinical testing, and highlight strategies our community can take to help stop a surge in new cases.
- It can help confirm current trends of COVID-19 infections in our community that are based on clinical data.
- It can increase confidence that clinical testing results are not biased by availability, time lags, and other factors.
SARS-CoV-2 Measurements in Wastewater
The plot shown above provides an overview of the results of SARS-CoV-2 measurements in wastewater over time. These are the results for two SARS-CoV-2 genes that are present in every variant: N gene and S gene. In addition, results for genes present in Omicron BA.2, BA.4 and BA.5 (LPPA24S) and Omicron BA.4 + BA.5 (HV69-70) are shown. The concentrations for these genes are normalized by an internal standard (copies of RNA from the genome of pepper mild mottle virus, PMMoV).
Alternate data formats
Stanford University is committed to providing an online environment that is accessible to everyone, including individuals with disabilities. If you cannot access the content on this page, please contact wastewater-dashboard@stanford.edu to obtain the data in alternate formats.